I am a passionate supporter of open science, contributing to many existing packages, as well as developing new libraries and tools as needed.
My own work is
- 100% open source
- 100% python
- 100% on github. Fork me!
My software span the following fields: machine learning, medical image processing, quality control, data visualization and research data management.
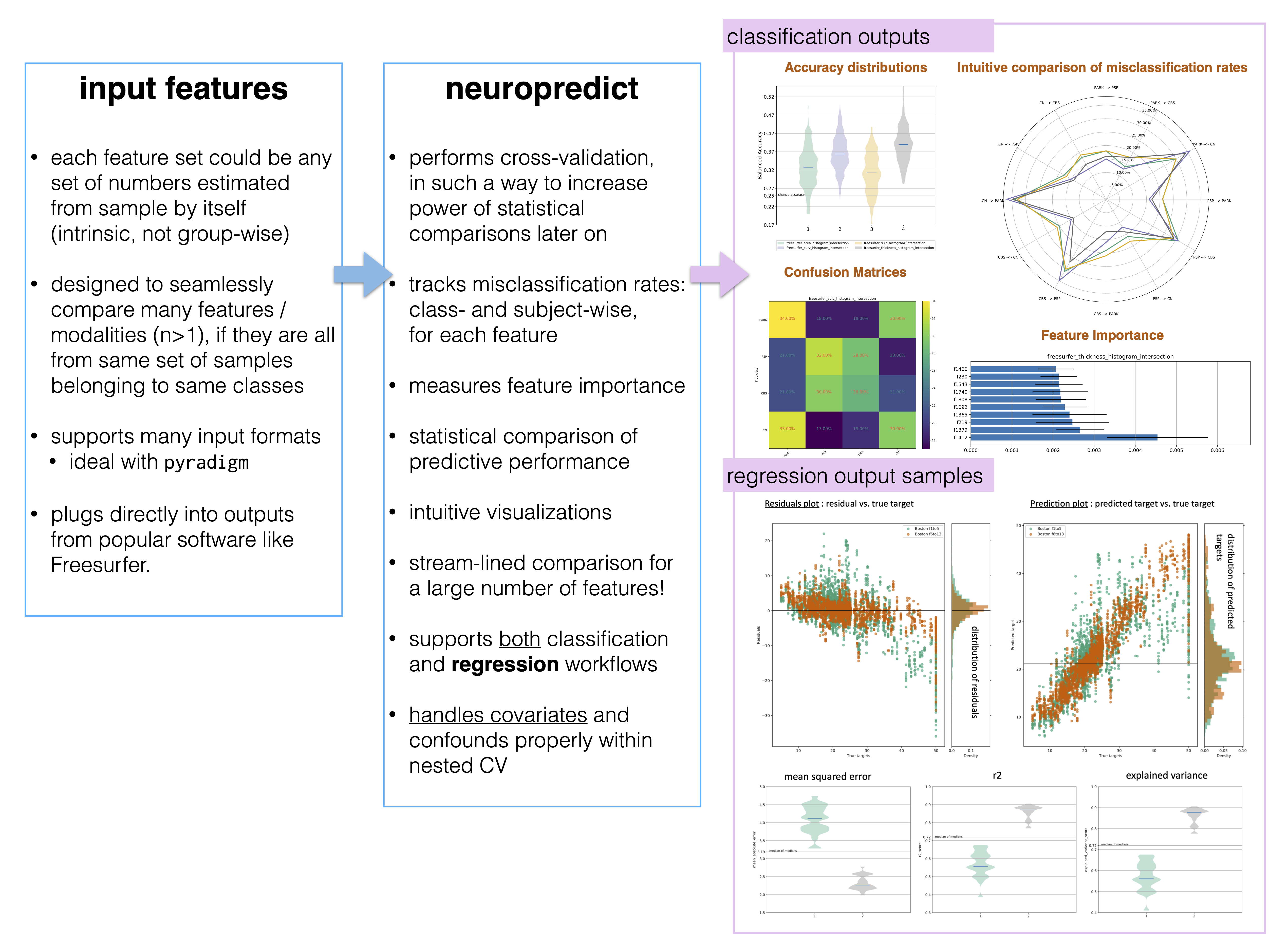
- neuropredict : easy and comprehensive analysis of predictive power of biomarkers, feature sets, following all the best practices. New release allowing prediction of continuous targets and regressing out covariates/confounds coming shortly.
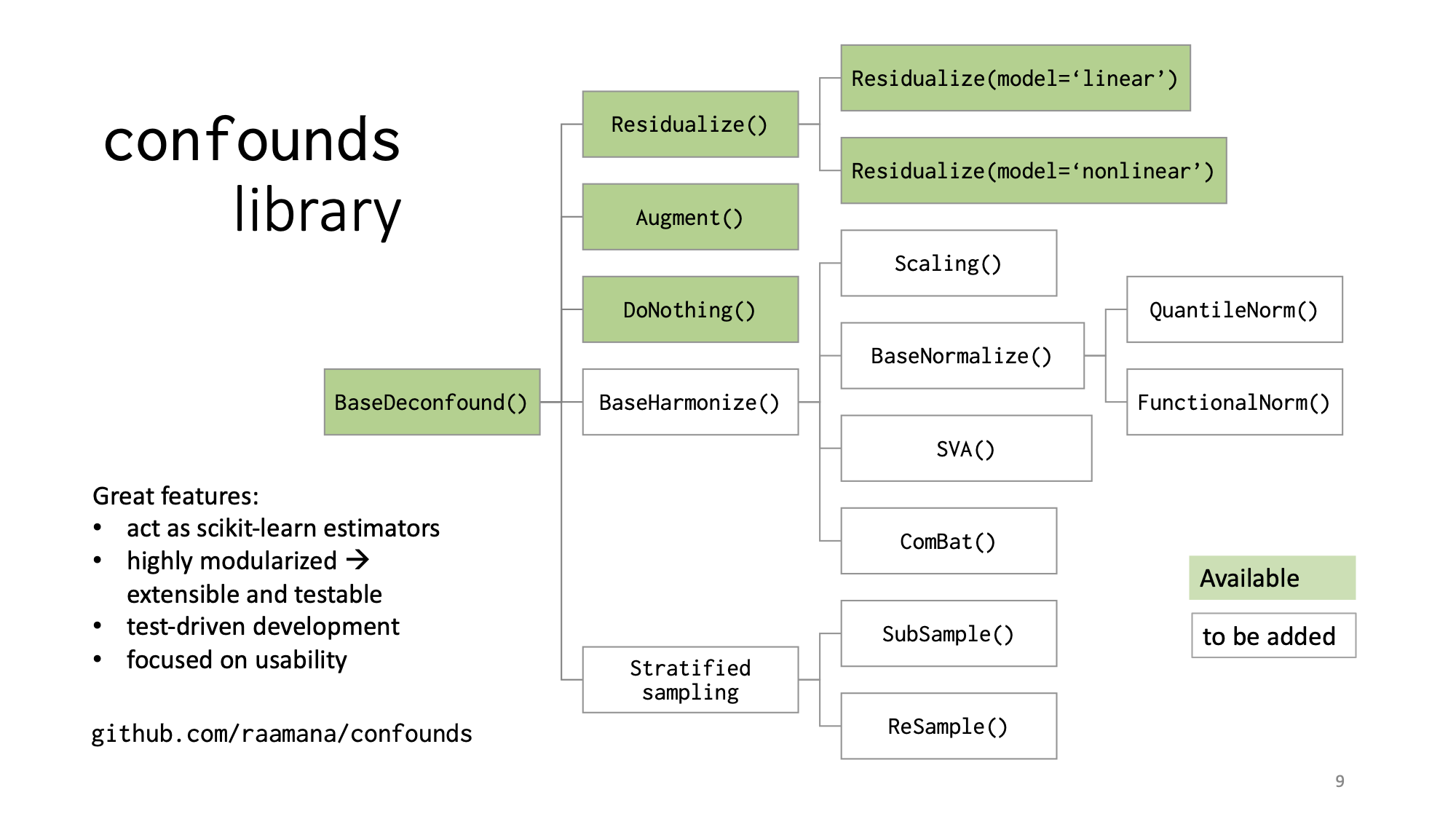
- confounds: python library implementing different deconfounding methods (such as Residualize / RegressOut, Augment etc) to help you conquer covariates/confounds in machine learning and neuroscience applications.
- visualqc: easy and rigorous quality control for neuroimaging data

- graynet : helps extract individualized single-subject networks (pairwise links between ROIs) from T1w MRI features such as cortical thickness, gray matter density, subcortical morphometric features, gyrification and curvature.

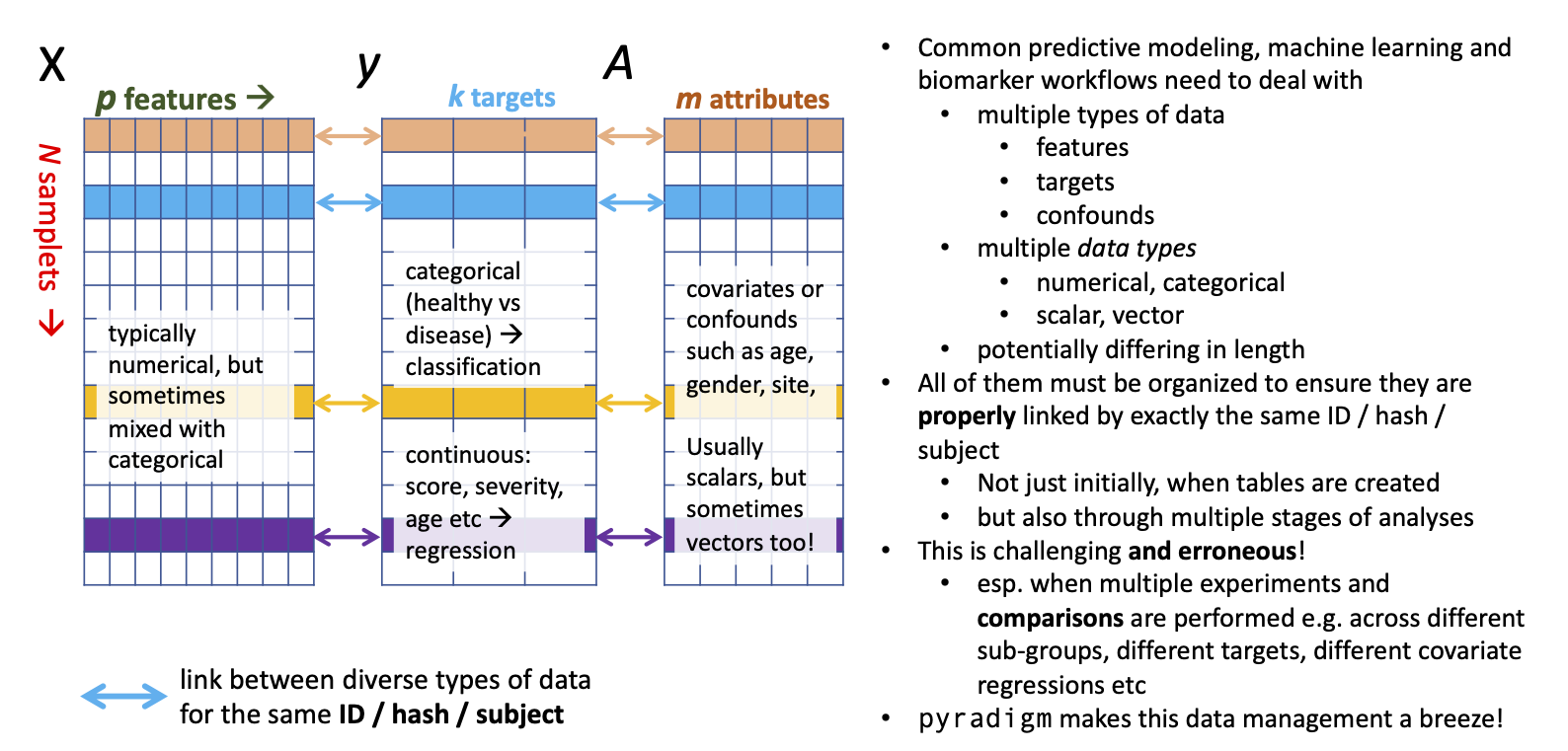
- Pyradigm: This is a Python class defining a machine learning dataset to ensure key-based correspondence within samples and maintaining integrity across samples. neuropredict leverages this package to a great effect. [repo]
- Kernel methods : Comprehensive kernel methods library for advanced machine learning applications in neuroscience [repo]

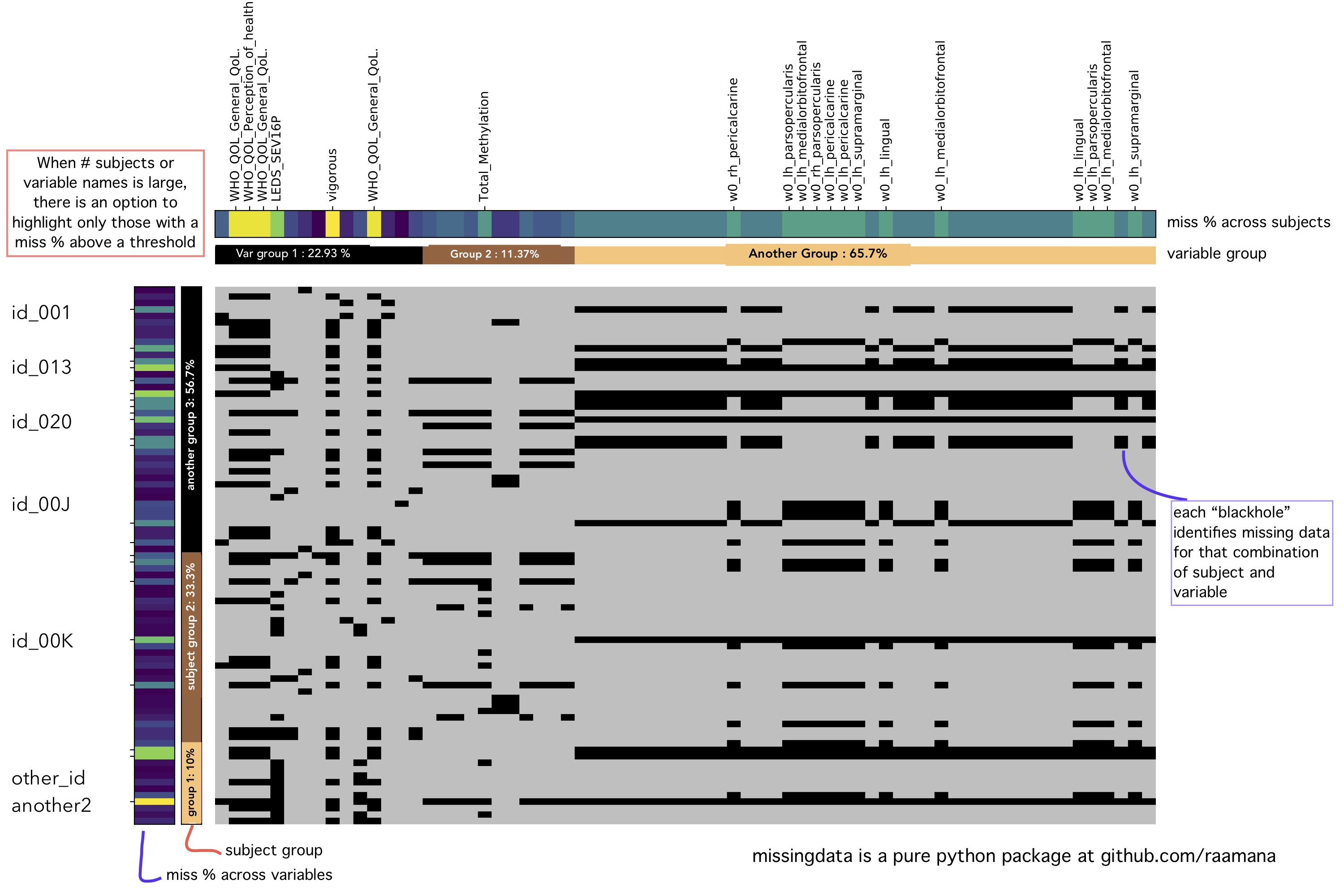
- Missing Data : visualization library for missing data, providing the comprehensive blackholes plot, summarizing the frequency of missingness across both variables as well as subjects. [repo]
- mrivis: visualization library to build advance medical image visualizations, such as the Carpet plot (to summarize 4D or higher-dimensional images, seen below for an fMRI scan), based on compositional classes like SlicePicker and Collage to enable building of heavily customized and advanced visualizations. [repo]

As well as variety of options to check the accuracy of alignment (or registration) between images from same modality (unimodal) or different modalities (cross-modal):
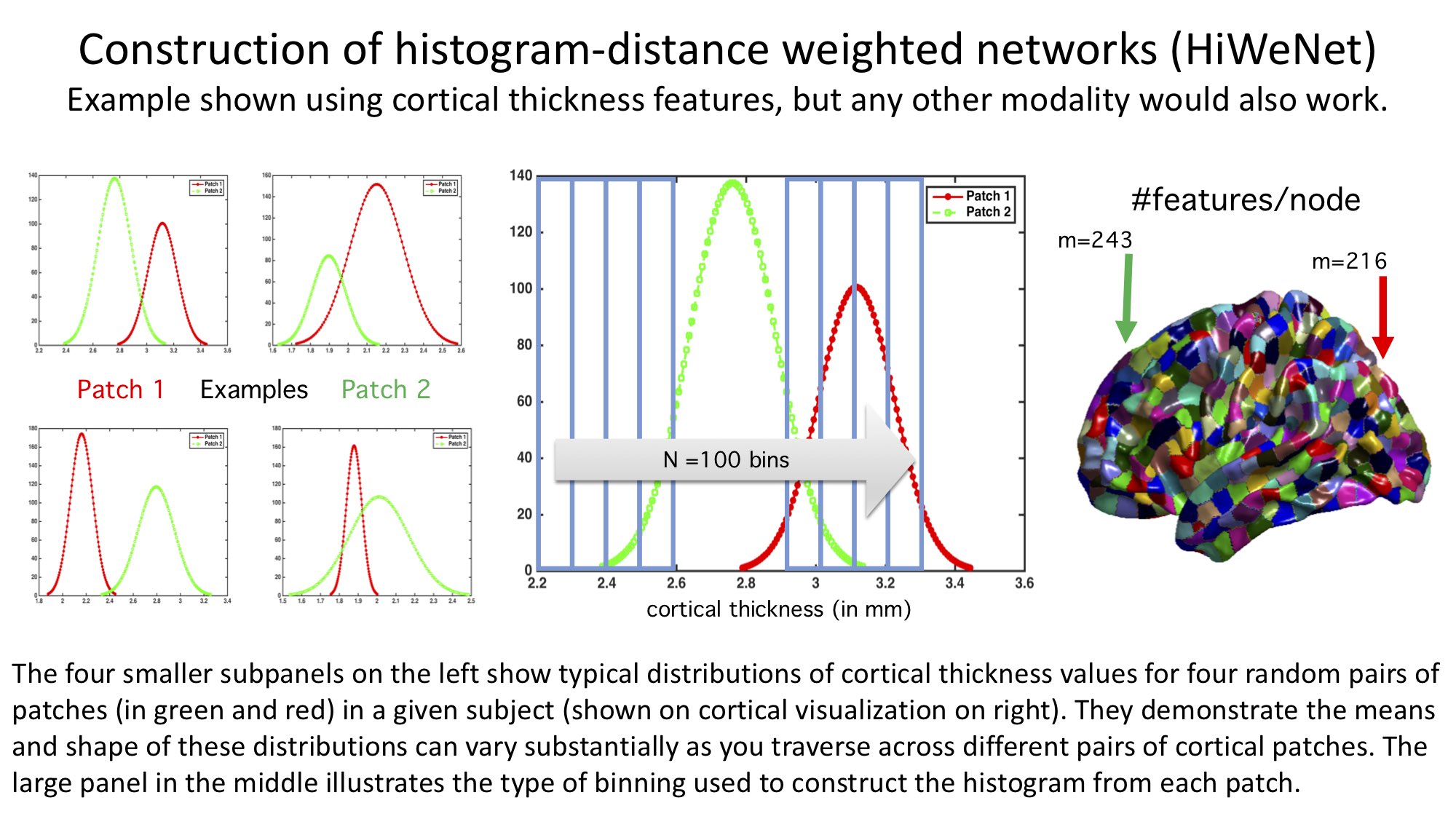
- hiwenet: Histogram-weighted Networks for Feature Extraction, Connectivity and Advanced Analysis in Neuroscience